c:ms:2026:lecture_note_week_04
This is an old revision of the document!
퀴즈 1 문제 중
모집단 Mean = 180; SD = 20, 정규분포일때
49. N=16의 샘플을 추출할 때 샘플들의 평균 분포가 갖는 표준편차 값은?
standard error 값을 묻는 질문이므로
se = sigma / sqrt(n) = 20 / 4 = 5
50. n=400일 때 샘플평균들의 분포가 갖는 표준편차 값은?
1
51. n=100 의 크기의 샘플을 취한다고 할 때 이 샘플의 평균값이 나올 구간을 99퍼센트의 확신성을 가지고 구하시오.
52. 위와합산
se값이 2이고 99퍼센트의 구간은 se값을 위로 3 밑은 3 포함하는 구간이 되므로 180-6, 180+6 이 정답
위의 문제는 모두 샘플평균들을 모아 놓은 집합의 평균과 표준편차를 (표준오차) 구하는 문제이다.
즉, 무 = 180, 시그마 = 20 일 때,
샘플평균들의 표준편차는 아래처럼 각각 5, 1, 2가 된다.
> rm(list=ls())
> rnorm2 <- function(n,mean,sd){
+ mean+sd*scale(rnorm(n))
+ }
> ss <- function(x) {
+ sum((x-mean(x))^2)
+ }
>
> mu = 180
> sigma = 20
> var = 400
>
> n.a <- 16
> n.b <- 400
> n.c <- 100
>
> se.a <- sigma/sqrt(n.a)
> se.b <- sigma/sqrt(n.b)
> se.c <- sigma/sqrt(n.c)
> se.a
[1] 5
> se.b
[1] 1
> se.c
[1] 2
Recap
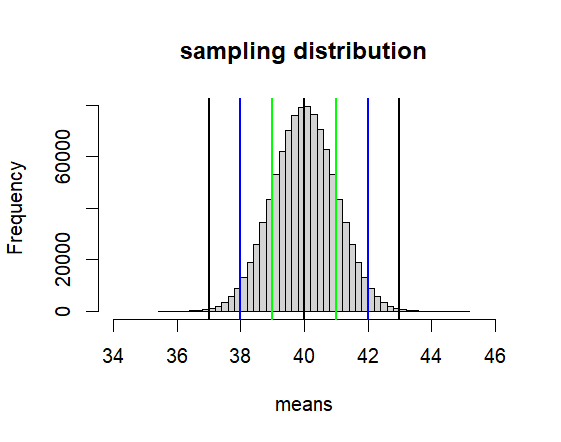
Distribution of Sample Means – mu = 40, sigma = 4 (hence var = 16) 인 모집단에서 n = n 사이즈의 샘플링을 무한 반복할 때 그 샘플평균들이 모인 집합
rscript01
rm(list=ls())
rnorm2 <- function(n,mean,sd){
mean+sd*scale(rnorm(n))
}
ss <- function(x) {
sum((x-mean(x))^2)
}
mu <- 40
sigma <- 4
iter <- 1000000
sz <- 16
se <- sigma/sqrt(16)
################################
means <- rnorm2(iter, mu, se)
hist(means, breaks=50,
xlim = c(mu-6*se, mu+6*se),
main = paste("sampling distribution"))
abline(v=mu, col='black', lwd=2)
lo1 <- mu - se*1
hi1 <- mu + se*1
lo2 <- mu - se*2
hi2 <- mu + se*2
lo3 <- mu - se*3
hi3 <- mu + se*3
abline(v=c(lo1, lo2, lo3, hi1, hi2, hi3),
col=c("green","blue", "black"),
lwd=2)
print(c(lo2, hi2))
m.samp <- 37
p.val <- pnorm(m.samp, mu, se)*2
p.val
z.cal <- (m.samp-mu)/se
z.cal
p.val <- pnorm(z.cal)*2
p.val
zmeans <- scale(means)
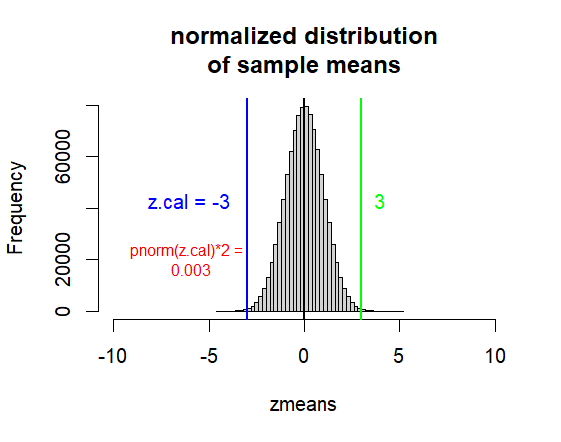
hist(zmeans, breaks=50,
xlim = c(0-10*1, 0+10*1),
main=("normalized distribution\nof sample means"))
abline(v=0, col="black", lwd=2)
abline(v=z.cal, col='blue', lwd=2)
abline(v=-z.cal, col="green", lwd=2)
text(x=-6, y=50000,
label=paste("z.cal =", z.cal),
pos = 1,
col="blue", cex=1)
text(x=4, y=50000,
label=paste(-z.cal),
pos=1,
col="green", cex=1)
text(x=-6, y=30000,
label=paste("pnorm(z.cal)*2 =", "\n",
round(p.val,3)),
pos = 1,
col="red", cex=.8)
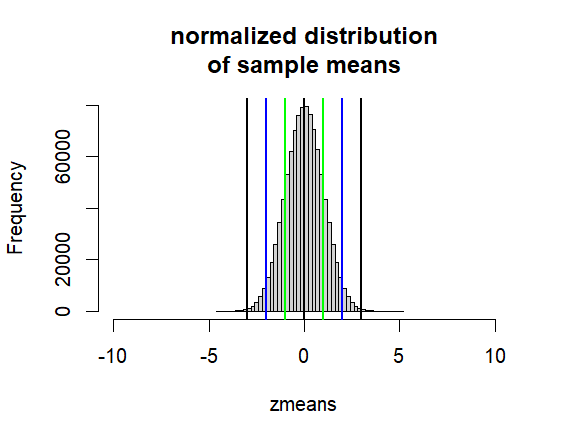
hist(zmeans, breaks=50,
xlim = c(0-10*1, 0+10*1),
main=("normalized distribution\nof sample means"))
abline(v=0, col="black", lwd=2)
abline(v=c(-1,-2,-3,1,2,3),
col=c("green", "blue", "black"), lwd=2)
z.cal
p.val
#####
# 위의 아이디어로는 z.cal 점수가
# +-2 밖에 있는지 보면 된다. 즉,
# 이는 prob가 0.05보다 작은지
# 보면 되는 것이다.
#####
# +-2 는 정확한 숫자가 아니고
# qnorm(.05/2) 에 해당하는 숫자
# 가 정확한 숫자
two.minus.exact <- qnorm(.05/2)
two.plus.exact <- qnorm(1-(.05/2))
c(two.minus.exact, two.plus.exact)
#####
# 그러나 R 사용시에는 z 점수로
# 판단하기 보다는
# 직접 구하는 prob.로 판단
pnorm(z.cal)*2
p.val
#####
# 위에서 그룹 간의 차이를
# standard error로 나누는 것에 주의
#
################
m.samp <- 43
sd.samp <- 4
sz <- 16
samp <- rnorm2(sz, m.samp, sd.samp)
diff <- m.samp - mu
se <- sd.samp / sqrt(sz)
t.cal <- diff/se
df <- sz-1
p.val <- pt(t.cal, df=df, lower.tail = F)*2
t.cal
df
p.val
t.test(samp, mu=mu)
#####
#
m.a <- 5.8
m.b <- 6.3
sd.a <- .5
sd.b <- .5
sz.a <- 16
sz.b <- 16
df.a <- sz.a-1
df.b <- sz.b-1
df <- df.a + df.b
a <- rnorm2(sz.a, m.a, sd.a)
b <- rnorm2(sz.b, m.b, sd.b)
diff <- m.a - m.b
pv <- (ss(a)+ss(b))/(df.a+df.b)
se <- sqrt(pv/sz.a+pv/sz.b)
t.cal <- diff / se
p.val <- pt(t.cal, df=df)*2
diff
se
t.cal
df
p.val
t.test(a,b, var.equal = T)
diff - se*2
diff + se*2
lo <- qt(.05/2,df)
lo
hi <- -lo
diff + se*lo
diff + se*hi
out01
> rm(list=ls())
> rnorm2 <- function(n,mean,sd){
+ mean+sd*scale(rnorm(n))
+ }
> ss <- function(x) {
+ sum((x-mean(x))^2)
+ }
>
> mu <- 40
> sigma <- 4
> iter <- 1000000
> sz <- 16
> se <- sigma/sqrt(16)
> ################################
> means <- rnorm2(iter, mu, se)
> hist(means, breaks=50,
+ xlim = c(mu-6*se, mu+6*se),
+ main = paste("sampling distribution"))
> abline(v=mu, col='black', lwd=2)
> lo1 <- mu - se*1
> hi1 <- mu + se*1
> lo2 <- mu - se*2
> hi2 <- mu + se*2
> lo3 <- mu - se*3
> hi3 <- mu + se*3
>
> abline(v=c(lo1, lo2, lo3, hi1, hi2, hi3),
+ col=c("green","blue", "black"),
+ lwd=2)
>
> print(c(lo2, hi2))
[1] 38 42
>
> m.samp <- 37
> p.val <- pnorm(m.samp, mu, se)*2
> p.val
[1] 0.002699796
> z.cal <- (m.samp-mu)/se
> z.cal
[1] -3
> p.val <- pnorm(z.cal)*2
> p.val
[1] 0.002699796
>
> zmeans <- scale(means)
> hist(zmeans, breaks=50,
+ xlim = c(0-10*1, 0+10*1),
+ main=("normalized distribution\nof sample means"))
> abline(v=0, col="black", lwd=2)
> abline(v=z.cal, col='blue', lwd=2)
> abline(v=-z.cal, col="green", lwd=2)
> text(x=-6, y=50000,
+ label=paste("z.cal =", z.cal),
+ pos = 1,
+ col="blue", cex=1)
> text(x=4, y=50000,
+ label=paste(-z.cal),
+ pos=1,
+ col="green", cex=1)
> text(x=-6, y=30000,
+ label=paste("pnorm(z.cal)*2 =", "\n",
+ round(p.val,3)),
+ pos = 1,
+ col="red", cex=.8)
>
> hist(zmeans, breaks=50,
+ xlim = c(0-10*1, 0+10*1),
+ main=("normalized distribution\nof sample means"))
> abline(v=0, col="black", lwd=2)
> abline(v=c(-1,-2,-3,1,2,3),
+ col=c("green", "blue", "black"), lwd=2)
>
> z.cal
[1] -3
> p.val
[1] 0.002699796
> #####
> # 위의 아이디어로는 z.cal 점수가
> # +-2 밖에 있는지 보면 된다. 즉,
> # 이는 prob가 0.05보다 작은지
> # 보면 되는 것이다.
> #####
> # +-2 는 정확한 숫자가 아니고
> # qnorm(.05/2) 에 해당하는 숫자
> # 가 정확한 숫자
> two.minus.exact <- qnorm(.05/2)
> two.plus.exact <- qnorm(1-(.05/2))
> c(two.minus.exact, two.plus.exact)
[1] -1.959964 1.959964
> #####
> # 그러나 R 사용시에는 z 점수로
> # 판단하기 보다는
> # 직접 구하는 prob.로 판단
> pnorm(z.cal)*2
[1] 0.002699796
> p.val
[1] 0.002699796
> #####
> # 위에서 그룹 간의 차이를
> # standard error로 나누는 것에 주의
> #
>
>
> ################
> m.samp <- 43
> sd.samp <- 4
> sz <- 16
> samp <- rnorm2(sz, m.samp, sd.samp)
> diff <- m.samp - mu
> se <- sd.samp / sqrt(sz)
> t.cal <- diff/se
> df <- sz-1
> p.val <- pt(t.cal, df=df, lower.tail = F)*2
> t.cal
[1] 3
> df
[1] 15
> p.val
[1] 0.008972737
> t.test(samp, mu=mu)
One Sample t-test
data: samp
t = 3, df = 15, p-value = 0.008973
alternative hypothesis: true mean is not equal to 40
95 percent confidence interval:
40.86855 45.13145
sample estimates:
mean of x
43
>
> #####
> #
> m.a <- 5.8
> m.b <- 6.3
> sd.a <- .5
> sd.b <- .5
> sz.a <- 16
> sz.b <- 16
> df.a <- sz.a-1
> df.b <- sz.b-1
> df <- df.a + df.b
> a <- rnorm2(sz.a, m.a, sd.a)
> b <- rnorm2(sz.b, m.b, sd.b)
> diff <- m.a - m.b
> pv <- (ss(a)+ss(b))/(df.a+df.b)
> se <- sqrt(pv/sz.a+pv/sz.b)
> t.cal <- diff / se
> p.val <- pt(t.cal, df=df)*2
>
> diff
[1] -0.5
> se
[1] 0.1767767
> t.cal
[1] -2.828427
> df
[1] 30
> p.val
[1] 0.008257336
> t.test(a,b, var.equal = T)
Two Sample t-test
data: a and b
t = -2.8284, df = 30, p-value = 0.008257
alternative hypothesis: true difference in means is not equal to 0
95 percent confidence interval:
-0.8610262 -0.1389738
sample estimates:
mean of x mean of y
5.8 6.3
> diff - se*2
[1] -0.8535534
> diff + se*2
[1] -0.1464466
> lo <- qt(.05/2,df)
> lo
[1] -2.042272
> hi <- -lo
> diff + se*lo
[1] -0.8610262
> diff + se*hi
[1] -0.1389738
>
rscript02
##### # m.a <- 5.8 m.b <- 6.3 sd.a <- .5 sd.b <- .5 sz.a <- 16 sz.b <- 16 df.a <- sz.a-1 df.b <- sz.b-1 df <- df.a + df.b a <- rnorm2(sz.a, m.a, sd.a) b <- rnorm2(sz.b, m.b, sd.b) diff <- m.a - m.b pv <- (ss(a)+ss(b))/(df.a+df.b) se <- sqrt(pv/sz.a+pv/sz.b) t.cal <- diff / se p.val <- pt(t.cal, df=df)*2 diff se t.cal df p.val t.test(a,b, var.equal = T) diff - se*2 diff + se*2 lo <- qt(.05/2,df) lo hi <- -lo diff + se*lo diff + se*hi ##### # t-test repeated measre ##### m.t1 <- 103 m.t2 <- 111 sd.t1 <- 10 sd.t2 <- 10 sz <- 160 t1 <- rnorm2(sz, m.t1, sd.t1) t2 <- rnorm2(sz, m.t2, sd.t2) t1 t2 mdiff <- m.t1-m.t2 diff <- t1-t2 sd.diff <- sd(diff) se <- sd.diff/sqrt(sz) t.cal <- mdiff/se p.val <- pt(t.cal, df=sz-1)*2 t.cal sz-1 p.val t.test(t1,t2, paired=T) two <- qt(.05/2, df=sz-1) two lo <- se*two hi <- -lo c(lo, hi) c(mdiff+lo, mdiff+hi)
rout02
>
>
>
>
> #####
> #
> m.a <- 5.8
> m.b <- 6.3
> sd.a <- .5
> sd.b <- .5
> sz.a <- 16
> sz.b <- 16
> df.a <- sz.a-1
> df.b <- sz.b-1
> df <- df.a + df.b
> a <- rnorm2(sz.a, m.a, sd.a)
> b <- rnorm2(sz.b, m.b, sd.b)
> diff <- m.a - m.b
> pv <- (ss(a)+ss(b))/(df.a+df.b)
> se <- sqrt(pv/sz.a+pv/sz.b)
> t.cal <- diff / se
> p.val <- pt(t.cal, df=df)*2
>
> diff
[1] -0.5
> se
[1] 0.1767767
> t.cal
[1] -2.828427
> df
[1] 30
> p.val
[1] 0.008257336
> t.test(a,b, var.equal = T)
Two Sample t-test
data: a and b
t = -2.8284, df = 30, p-value = 0.008257
alternative hypothesis: true difference in means is not equal to 0
95 percent confidence interval:
-0.8610262 -0.1389738
sample estimates:
mean of x mean of y
5.8 6.3
> diff - se*2
[1] -0.8535534
> diff + se*2
[1] -0.1464466
> lo <- qt(.05/2,df)
> lo
[1] -2.042272
> hi <- -lo
> diff + se*lo
[1] -0.8610262
> diff + se*hi
[1] -0.1389738
>
> #####
> # t-test repeated measre
> #####
> m.t1 <- 103
> m.t2 <- 111
> sd.t1 <- 10
> sd.t2 <- 10
> sz <- 160
> t1 <- rnorm2(sz, m.t1, sd.t1)
> t2 <- rnorm2(sz, m.t2, sd.t2)
> t1
[,1]
[1,] 111.36104
[2,] 88.01731
[3,] 101.88320
[4,] 103.10436
[5,] 96.38094
[6,] 110.28539
[7,] 98.80045
[8,] 86.31780
[9,] 128.73687
[10,] 102.23723
[11,] 94.90932
[12,] 110.74820
[13,] 108.90433
[14,] 91.11044
[15,] 83.69413
[16,] 84.12565
[17,] 110.63493
[18,] 77.88247
[19,] 114.62381
[20,] 109.08372
[21,] 105.43251
[22,] 93.28446
[23,] 112.45340
[24,] 108.82774
[25,] 101.87046
[26,] 99.06745
[27,] 105.41497
[28,] 89.35983
[29,] 128.33953
[30,] 101.82933
[31,] 84.43396
[32,] 107.31985
[33,] 94.50306
[34,] 98.51383
[35,] 100.37196
[36,] 104.39854
[37,] 99.00553
[38,] 100.39100
[39,] 98.94237
[40,] 94.04721
[41,] 91.55691
[42,] 77.02969
[43,] 100.65928
[44,] 99.50989
[45,] 113.40564
[46,] 91.27212
[47,] 96.54430
[48,] 103.67181
[49,] 91.91200
[50,] 95.86468
[51,] 97.73431
[52,] 105.95878
[53,] 99.40692
[54,] 114.89231
[55,] 110.23953
[56,] 110.65776
[57,] 95.35294
[58,] 114.74190
[59,] 107.10249
[60,] 97.93327
[61,] 114.29149
[62,] 106.77413
[63,] 89.85116
[64,] 100.92937
[65,] 110.57659
[66,] 118.43433
[67,] 97.26787
[68,] 112.06303
[69,] 101.08834
[70,] 112.54527
[71,] 103.74242
[72,] 107.31976
[73,] 114.14557
[74,] 96.41347
[75,] 96.73140
[76,] 92.48801
[77,] 93.13216
[78,] 93.39353
[79,] 106.83687
[80,] 95.43550
[81,] 99.92717
[82,] 105.47433
[83,] 88.13565
[84,] 104.37033
[85,] 96.23481
[86,] 105.73652
[87,] 99.62358
[88,] 112.79561
[89,] 111.78083
[90,] 114.73846
[91,] 98.61353
[92,] 121.41442
[93,] 104.81865
[94,] 100.92946
[95,] 107.41369
[96,] 98.22645
[97,] 104.94036
[98,] 93.38986
[99,] 107.18154
[100,] 108.80844
[101,] 117.97939
[102,] 103.40657
[103,] 99.54187
[104,] 98.22691
[105,] 99.13327
[106,] 93.54839
[107,] 99.47141
[108,] 72.82718
[109,] 120.41493
[110,] 106.81977
[111,] 104.10554
[112,] 92.11256
[113,] 117.84020
[114,] 106.80209
[115,] 123.11219
[116,] 112.60503
[117,] 113.01015
[118,] 95.06184
[119,] 97.10124
[120,] 88.02648
[121,] 103.98118
[122,] 112.38688
[123,] 100.76566
[124,] 104.56130
[125,] 110.20566
[126,] 108.55945
[127,] 101.47467
[128,] 100.21853
[129,] 103.10659
[130,] 95.19338
[131,] 98.03036
[132,] 107.44486
[133,] 100.49136
[134,] 105.64403
[135,] 103.33323
[136,] 111.37567
[137,] 88.13074
[138,] 106.90384
[139,] 100.01857
[140,] 110.50553
[141,] 124.36441
[142,] 106.98552
[143,] 115.77759
[144,] 101.10420
[145,] 105.26656
[146,] 93.98217
[147,] 120.60988
[148,] 94.68497
[149,] 127.71822
[150,] 128.63994
[151,] 106.18538
[152,] 92.98331
[153,] 99.14643
[154,] 110.37932
[155,] 104.60248
[156,] 106.81372
[157,] 94.45348
[158,] 113.53202
[159,] 107.81640
[160,] 87.36641
attr(,"scaled:center")
[1] 0.02797819
attr(,"scaled:scale")
[1] 1.028345
> t2
[,1]
[1,] 104.52214
[2,] 106.01539
[3,] 118.86196
[4,] 110.72811
[5,] 121.76890
[6,] 104.08372
[7,] 104.79700
[8,] 130.54567
[9,] 104.51772
[10,] 98.46445
[11,] 102.79140
[12,] 120.40580
[13,] 112.91734
[14,] 113.77920
[15,] 114.20003
[16,] 107.49174
[17,] 101.19277
[18,] 115.36843
[19,] 119.91569
[20,] 106.06605
[21,] 123.11635
[22,] 93.79225
[23,] 93.48746
[24,] 117.74609
[25,] 109.58166
[26,] 134.83143
[27,] 98.45053
[28,] 106.25705
[29,] 100.76346
[30,] 117.52330
[31,] 99.08305
[32,] 120.38723
[33,] 116.10505
[34,] 120.00785
[35,] 116.23227
[36,] 116.00613
[37,] 124.99957
[38,] 115.16024
[39,] 114.95141
[40,] 98.03156
[41,] 109.35921
[42,] 108.94960
[43,] 106.56490
[44,] 116.28102
[45,] 116.59853
[46,] 108.34954
[47,] 113.88005
[48,] 89.61658
[49,] 106.68461
[50,] 124.51694
[51,] 124.60305
[52,] 103.93134
[53,] 118.46683
[54,] 118.84622
[55,] 118.55730
[56,] 120.96029
[57,] 120.91002
[58,] 93.65926
[59,] 118.82763
[60,] 113.24234
[61,] 113.75956
[62,] 111.12494
[63,] 107.98393
[64,] 118.47903
[65,] 108.81494
[66,] 122.69894
[67,] 108.42655
[68,] 130.67077
[69,] 97.24069
[70,] 110.17917
[71,] 103.99463
[72,] 125.31486
[73,] 91.26600
[74,] 105.84776
[75,] 123.22794
[76,] 110.03864
[77,] 115.89615
[78,] 112.08779
[79,] 112.91045
[80,] 120.60075
[81,] 105.78417
[82,] 92.46600
[83,] 88.53759
[84,] 127.18477
[85,] 122.35360
[86,] 123.61826
[87,] 110.69036
[88,] 110.69824
[89,] 107.37120
[90,] 107.72845
[91,] 112.04867
[92,] 96.05635
[93,] 108.67360
[94,] 118.85047
[95,] 103.94559
[96,] 110.11686
[97,] 122.59443
[98,] 110.32224
[99,] 105.05468
[100,] 119.83378
[101,] 116.87027
[102,] 113.08225
[103,] 109.16396
[104,] 108.50073
[105,] 105.41310
[106,] 95.32322
[107,] 105.61400
[108,] 129.13587
[109,] 86.96069
[110,] 115.11906
[111,] 106.76034
[112,] 102.69395
[113,] 97.25614
[114,] 105.64480
[115,] 111.23453
[116,] 117.35671
[117,] 120.65920
[118,] 108.53758
[119,] 109.18312
[120,] 112.51380
[121,] 119.76297
[122,] 132.25435
[123,] 118.96046
[124,] 108.72113
[125,] 124.70416
[126,] 91.89676
[127,] 117.43380
[128,] 116.62698
[129,] 94.52859
[130,] 106.74281
[131,] 109.20631
[132,] 112.79079
[133,] 113.50772
[134,] 91.10812
[135,] 77.92339
[136,] 118.20013
[137,] 106.33092
[138,] 112.72908
[139,] 110.12456
[140,] 114.61151
[141,] 123.38285
[142,] 119.22545
[143,] 116.54886
[144,] 111.95272
[145,] 112.92325
[146,] 104.37119
[147,] 110.30857
[148,] 103.80554
[149,] 131.22599
[150,] 107.36467
[151,] 110.81009
[152,] 95.49339
[153,] 107.73582
[154,] 105.47526
[155,] 119.48971
[156,] 114.18188
[157,] 93.91213
[158,] 107.63677
[159,] 125.62210
[160,] 101.10034
attr(,"scaled:center")
[1] -0.03441822
attr(,"scaled:scale")
[1] 0.9901942
> mdiff <- m.t1-m.t2
> diff <- t1-t2
> sd.diff <- sd(diff)
> se <- sd.diff/sqrt(sz)
> t.cal <- mdiff/se
> p.val <- pt(t.cal, df=sz-1)*2
> t.cal
[1] -7.249999
> sz-1
[1] 159
> p.val
[1] 1.713273e-11
> t.test(t1,t2, paired=T)
Paired t-test
data: t1 and t2
t = -7.25, df = 159, p-value = 1.713e-11
alternative hypothesis: true mean difference is not equal to 0
95 percent confidence interval:
-10.179307 -5.820693
sample estimates:
mean difference
-8
> two <- qt(.05/2, df=sz-1)
> two
[1] -1.974996
> lo <- se*two
> hi <- -lo
> c(lo, hi)
[1] -2.179307 2.179307
> c(mdiff+lo, mdiff+hi)
[1] -10.179307 -5.820693
>
c/ms/2026/lecture_note_week_04.1774974886.txt.gz · Last modified: by hkimscil