rs.hypothesis.testing
rm(list=ls())
rnorm2 <- function(n,mean,sd){ mean+sd*scale(rnorm(n)) }
ss <- function(x) { sum((x-mean(x))^2) }
z.test <- function(sam, mu, sigma) {
diff <-mean(sam) - mu
se <- sigma / sqrt(length(sam))
z.cal <- diff / se
p.val <- pnorm(abs(z.cal), lower.tail = F)*2
two <- abs(qnorm(.05/2))
return(cat(
" z value:", round(z.cal,5), '\n',
"p value:", round(p.val,8), '\n',
"diff: ", mean(sam), "-", mu, "=", mean(sam)-mu, '\n',
"se: ", se, '\n',
"95% CI: ", c(mu-two*se, mu+two*se) ) )
}
################################
set.seed(1001)
N.p <- 1000000
m.p <- 100
sd.p <- 10
p1 <- rnorm2(N.p, m.p, sd.p)
p2 <- rnorm2(N.p, m.p+6, sd.p)
################################
sz <- 10
iter <- 100000
# n = 10 일 때의 p1에 대한 sampling dist 은 아래 시뮬레이션으로
# 구해볼 수 있다.
means <- rep(NA, iter)
for (i in 1:iter) {
s1 <- sample(p1, sz, replace = T)
means[i] <- mean(s1)
}
mean(means)
var(means)
sd(means)
# CLT에 의하면 위이 값은
m.means <- mean(p1)
ms.means <- c(var(p1)/sz)
sd.means <- c(sqrt(var(p1)/sz))
m.means
ms.means
sd.means
# 위의 시뮬레이션으로 구한 sampling dist 대신에
# 정확한 mean과 se 값을 갖는 집합 sdc를 만든다
sdc <- rnorm2(iter, m.means, sd.means)
mean(sdc)
var(sdc)
sd(sdc)
zsdc <- scale(sdc)
m.zsdc <- mean(zsdc)
ms.zsdc <- var(zsdc)
sd.zsdc <- sd(zsdc)
m.zsdc
ms.zsdc
sd.zsdc
se2 <- c(sqrt(var(p2)/sz))
z.p2 <- c((mean(p2)-mean(p1))/se2)
sdc2 <- rnorm2(iter, mean(p2), se2)
zsdc2 <- scale(sdc2)+z.p2
mean(zsdc2)
sd(zsdc2)
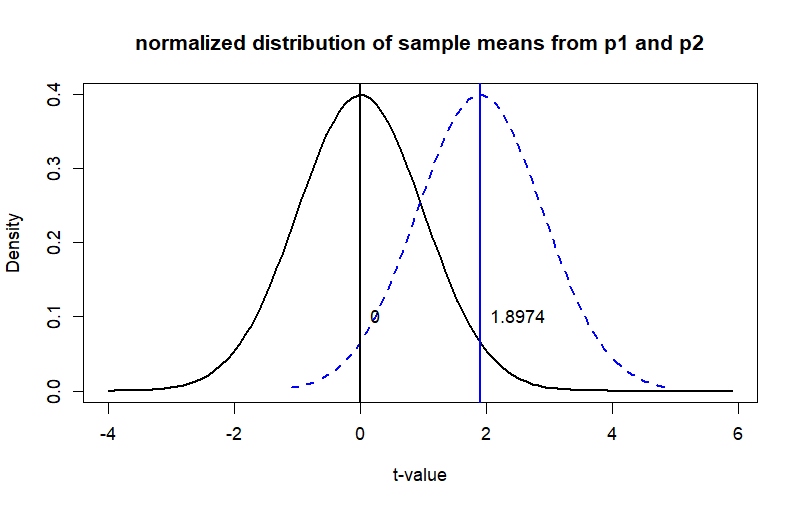
curve(dnorm(x), from = -4, to = z.p2+4,
main = "normalized distribution of sample means from p1 and p2",
ylab = "Density", xlab = "t-value", col = "black", lwd = 2)
curve(dnorm(x-(z.p2)), from = z.p2-3, to = z.p2+3, add = T,
main = "Distribution Curve",
ylab = "Density", xlab = "t-value", col = "blue", lwd = 2, lty=2)
abline(v=mean(zsdc), col='black', lwd=2)
abline(v=mean(zsdc2), col='blue', lwd=2)
mean(zsdc2)
text(x=mean(zsdc), y=.1, label=paste(round(mean(zsdc),4)), pos=4)
text(x=mean(zsdc2), y=.1, label=paste(round(mean(zsdc2),4)), pos=4)
#
lo1 <- qnorm(.32/2)
hi1 <- -lo1
lo2 <- qnorm(.05/2)
hi2 <- -lo2
lo3 <- qnorm(.01/2)
hi3 <- -lo3
c(lo1,hi1)
c(lo2,hi2)
c(lo3,hi3)
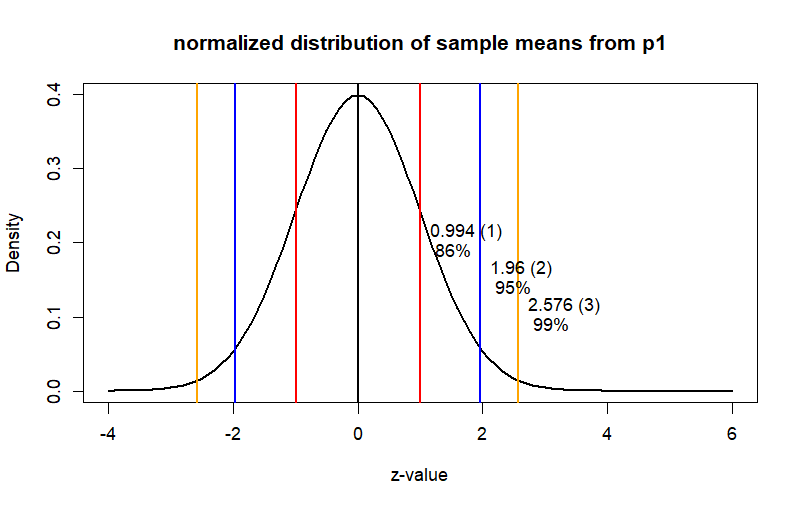
curve(dnorm(x), from = -4, to = 2+4,
main = "normalized distribution of sample means from p1",
ylab = "Density", xlab = "z-value", col = "black", lwd = 2)
abline(v=0, col="black", lwd=2)
abline(v=c(lo1, hi1, lo2, hi2, lo3, hi3),
col=c("red","red", "blue", "blue", "orange", "orange"),
lwd=2)
text(x=hi1, y=.2, label=paste(round(hi1,3), "(1)", "\n","86%"), pos=4)
text(x=hi2, y=.15, label=paste(round(hi2,3),"(2)", "\n","95%"), pos=4)
text(x=hi3, y=.1, label=paste(round(hi3,3), "(3)", "\n","99%"), pos=4)
mean.of.sample.a <- mean(sdc)+ 1.5*sd(sdc)
mean.of.sample.a
diff <- (mean.of.sample.a - mean(sdc))
se.z <- sd(p1)/sqrt(sz)
diff
se.z
z.score <- diff / se.z
z.score
prob <- pnorm(z.score, lower.tail = F)*2
prob
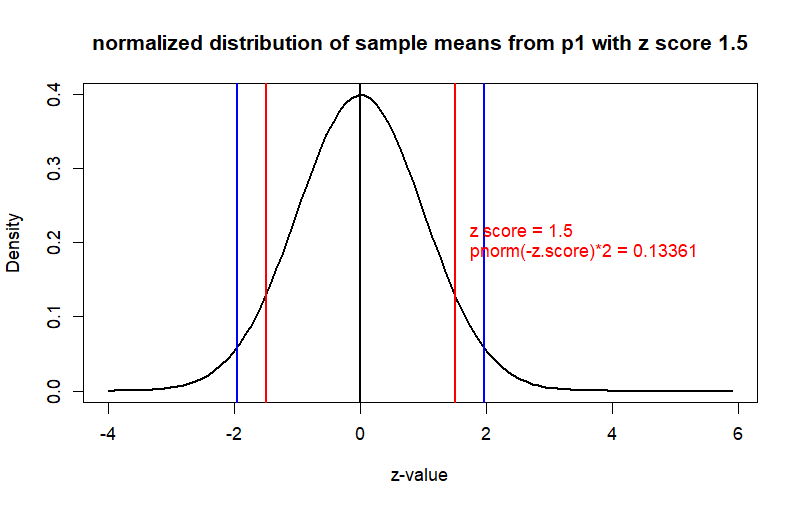
curve(dnorm(x), from = -4, to = z.p2+4,
main = "normalized distribution of sample means from p1 with z score 1.5",
ylab = "Density", xlab = "z-value", col = "black", lwd = 2)
abline(v=0, col="black", lwd=2)
abline(v=c(lo2, hi2), col=c("blue"), lwd=2)
abline(v=z.score,col='red', lwd=2)
abline(v=-z.score, col='red', lwd=2)
text(x=z.score, y=0.2,
label = paste(" z score =", z.score,
"\n", "pnorm(-z.score)*2 =", round(prob,5)),
pos=4, col='red')
# 새로운 UI로 게임을 하도록 한 후
# UI점수를 10명에게 구했다고 가정하고
# 새로운 UI점수가 기존의 p1 paramter와
# 다른지 테스트 해보라
# z-test 가설검증에 실패하는 경우, 사실은 성공해야 하는데
# p2의 평균이 106점이고 p1은 100점이니 p2에서 sample을 취
# 하면 샘플의 평균과 p1의 평균은 다르다고 판단될 것이다.
# 아래는 그럼에도 불구하고 실패하는 경우이다.
set.seed(111)
smp <- sample(p2, sz, replace=T)
m.smp <- mean(smp)
m.smp
diff <- m.smp - mean(p1)
se.z <- sqrt(var(p1)/sz)
z.cal1 <- diff / se.z
prob1 <- pnorm(abs(z.cal1), lower.tail = F)*2
print(c(z.cal1, sz, prob1))
z.test(smp, mean(p1), sd(p1))
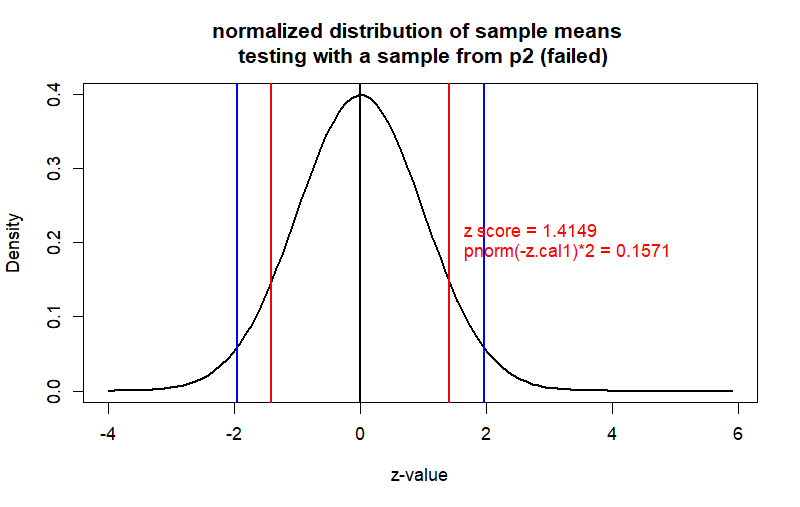
curve(dnorm(x), from = -4, to = z.p2+4,
main = "normalized distribution of sample means \n testing with a sample from p2 (failed)",
ylab = "Density", xlab = "z-value", col = "black", lwd = 2)
abline(v=0, col="black", lwd=2)
abline(v=c(lo2, hi2), col=c("blue"), lwd=2)
abline(v=z.cal1,col='red', lwd=2)
abline(v=-z.cal1, col='red', lwd=2)
text(x=z.cal1, y=0.2,
label = paste(" z score =", round(z.cal1,4),
"\n", "pnorm(-z.cal1)*2 =", round(prob1,4)),
pos=4, col='red')
# 같은 방법으로 했는데 성공한 경우
set.seed(211)
smp <- sample(p2,sz,replace=T)
m.smp <- mean(smp)
m.smp
diff <- m.smp - mean(p1)
se.z <- sqrt(var(p1)/sz)
z.cal2 <- diff / se.z
prob2 <- pnorm(abs(z.cal2), lower.tail = F)*2
print(c(z.cal2, sz, prob2))
z.test(smp, mean(p1), sd(p1))
z.p2 <- (mean(p2)-mean(p1))/se2
z.p2
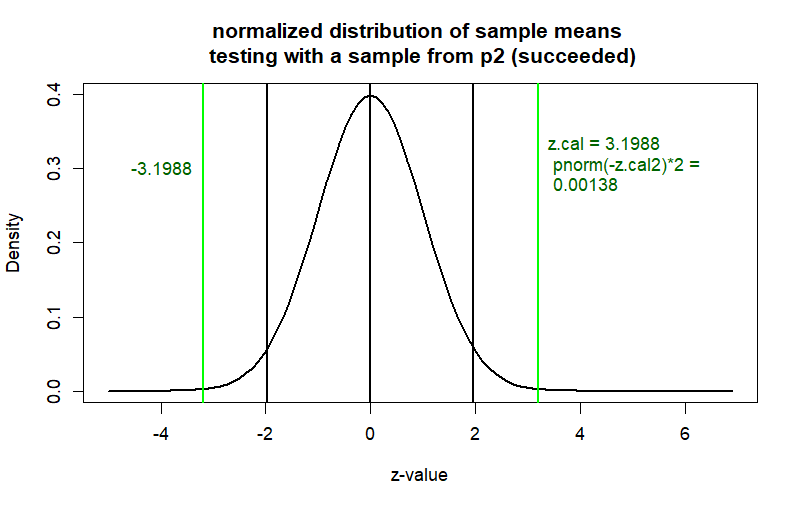
curve(dnorm(x), from = -5, to = z.p2+5,
main = "normalized distribution of sample means \n testing with a sample from p2 (succeeded)",
ylab = "Density", xlab = "z-value", col = "black", lwd = 2)
abline(v=0, col='black', lwd=2)
z.cal1
z.cal2
two <- qnorm(.05/2)
two
abline(v=c(two, -two), col='black', lwd=2)
abline(v=c(-z.cal2, z.cal2), col='green', lwd=2)
text(x=z.cal2, y=.3,
label=paste("z.cal =", round(z.cal2,4), "\n",
"pnorm(-z.cal2)*2 =", "\n", round(prob2,5)),
col="darkgreen", cex=1, pos=4)
text(x=-z.cal2, y=.3,
label=paste(round(-z.cal2,4)),
col="darkgreen", cex=1, pos=2)
# type i and type ii error
z.p2 <- (mean(p2)-mean(p1))/se2
z.p2
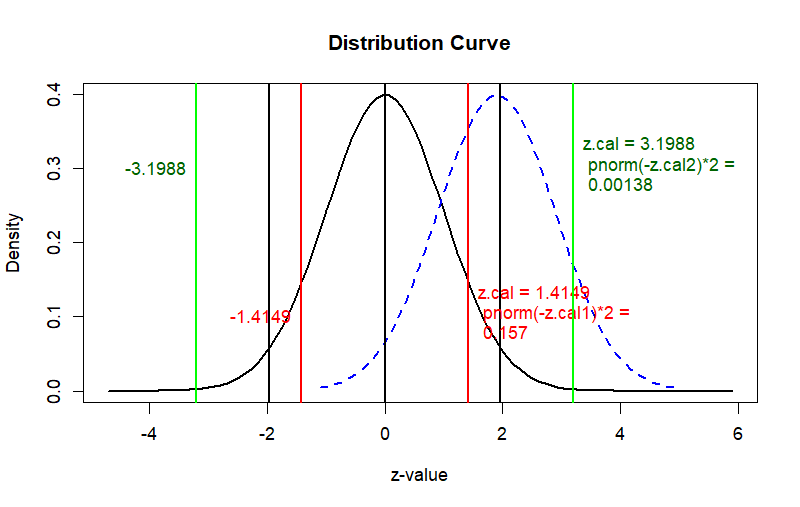
curve(dnorm(x), from = -4.7, to = z.p2+4,
main = "Distribution Curve",
ylab = "Density", xlab = "z-value", col = "black", lwd = 2)
curve(dnorm(x-(z.p2)), from = z.p2-3, to = z.p2+3, add = T,
main = "Distribution Curve",
ylab = "Density", xlab = "z-value", col = "blue", lwd = 2, lty=2)
abline(v=0, col='black', lwd=2)
z.cal1
z.cal2
two <- qnorm(.05/2)
two
abline(v=c(two, -two), col='black', lwd=2)
abline(v=c(-z.cal1, z.cal1), col='red', lwd=2)
text(x=z.cal1, y=.1,
label=paste("z.cal =", round(z.cal1,4), "\n",
"pnorm(-z.cal1)*2 =", "\n", round(prob1,3)),
col="red", cex=1, pos=4)
text(x=-z.cal1, y=.1,
label=paste(round(-z.cal1,4)),
col="red", cex=1, pos=2)
abline(v=c(-z.cal2, z.cal2), col='green', lwd=2)
text(x=z.cal2, y=.3,
label=paste("z.cal =", round(z.cal2,4), "\n",
"pnorm(-z.cal2)*2 =", "\n", round(prob2,5)),
col="darkgreen", cex=1, pos=4)
text(x=-z.cal2, y=.3,
label=paste(round(-z.cal2,4)),
col="darkgreen", cex=1, pos=2)
############################
# one sample t-test
############################
set.seed(99)
sz <- 20
smp <- sample(p2, sz, replace = T)
m.smp <- mean(smp)
diff <- m.smp - mean(p1)
se.z <- sqrt(var(smp)/sz)
df.smp <- sz - 1
t.cal <- (diff/se.z)
prob <- pt(t.cal, df.smp, lower.tail = F)*2
se.z
qt(.05/2, df.smp)
lo2 <- qt(.05/2, df.smp)
hi2 <- -lo2
m.smp+lo2*se.z
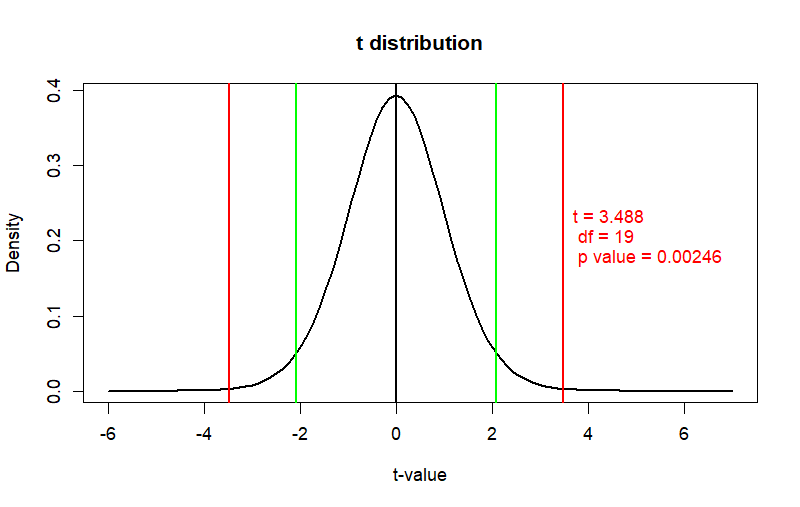
curve(dt(x, df.smp), from = -6, to = 7,
main = "t distribution",
ylab = "Density", xlab = "t-value", col = "black", lwd = 2)
abline(v=0, col="black", lwd=2)
abline(v=c(lo2, hi2), col="green", lwd=2)
abline(v=c(-t.cal, t.cal), col="red", lwd=2)
text(x=t.cal,y=.2,
label=paste("t =", round(t.cal,3),
"\n", "df =", df.smp,
"\n", "p value =", round(prob, 5)),
pos = 4, col="red", cex=1)
prob
print(c(t.cal, df.smp, prob))
print(c(m.smp+lo2*se.z, m.smp+hi2*se.z))
cat("t =", t.cal, ", df =", round(df.smp,0), ", p-value =", prob,
"\n", "95% confidence interval =", m.smp+lo2*se.z, m.smp+hi2*se.z)
t.test(smp, mu=mean(p1))
#################################
# t-test 2 group
#################################
set.seed(169)
sz.a <- 25
sz.b <- 25
group.a <- sample(p1, sz.a)
group.b <- sample(p2, sz.b)
m.a <- mean(group.a)
m.b <- mean(group.b)
ss.a <- ss(group.a)
ss.b <- ss(group.b)
df.a <- sz.a - 1
df.b <- sz.b - 1
df <- df.a+df.b
ss.a
ss.b
df.a
df.b
pooled.v <- (ss.a+ss.b)/(df.a+df.b)
pooled.v
se.s <- sqrt(pooled.v/sz.a+pooled.v/sz.b)
se.s
diff <- m.a-m.b
t.cal <- diff/se.s
t.cal
prob <- pt(abs(t.cal), df=df, lower.tail = F)*2
t.cal
df
prob
t.test(group.a, group.b, var.equal = T)
lo2 <- qt(.05/2, df)
hi2 <- -lo2
c(lo2, hi2)
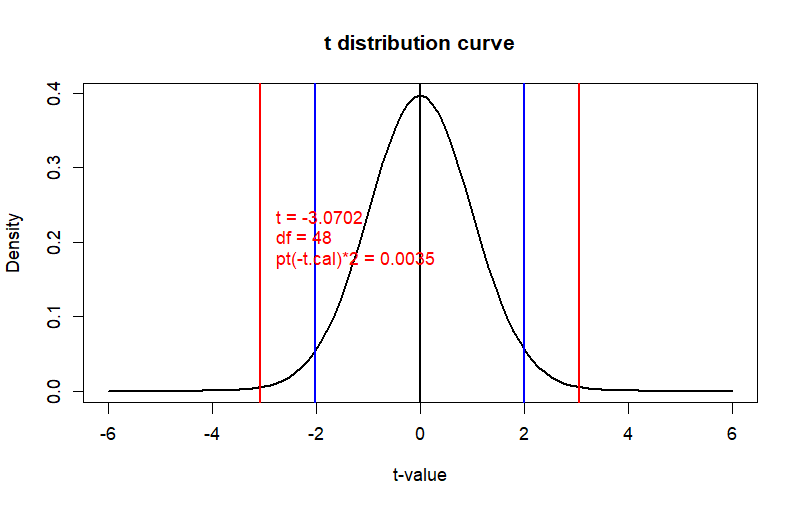
curve(dt(x, df=df), from = -6, to = 6,
main = "t distribution curve",
ylab = "Density", xlab = "t-value", col = "black", lwd = 2)
abline(v=0, col="black", lwd=2)
abline(v=c(lo2, hi2), col=c("blue"), lwd=2)
abline(v=c(t.cal, -t.cal), col='red', lwd=2)
text(x=t.cal, y=0.2,
label = paste(" t =", round(t.cal, 4),
"\n",
"df =", df, "\n",
"pt(-t.cal)*2 =", round(prob, 4)),
pos=4, col='red')
print(paste(t.cal, df, prob))
t.test(group.a, group.b, var.equal = T)
t.cal
# t.cal=diff/se
t.cal * se.s
diff
diff+lo2*se.s
diff+hi2*se.s
(t.cal+lo2)*se.s
(t.cal+hi2)*se.s
######################
# 4번째 케이스 t-test
######################
set.seed(37)
sz <- 40
time.a <- sample(p1,sz)
time.b <- sample(p2,sz)
diff.time <- time.a-time.b
m.a <- mean(time.a)
m.b <- mean(time.b)
diff <- m.a-m.b
diff
se.s <- sd(diff.time)/sqrt(sz)
t.cal <- diff/se.s
df <- sz - 1
prob <- pt(abs(t.cal), df=sz-1, lower.tail = F)*2
t.cal
df
prob
diff
m.diff.time <- mean(diff.time)
se.s
t.test(time.a, time.b, paired=T)
m.diff.time
se.s
lo1 <- qt(0.32/2,sz-1)
hi1 <- -lo1
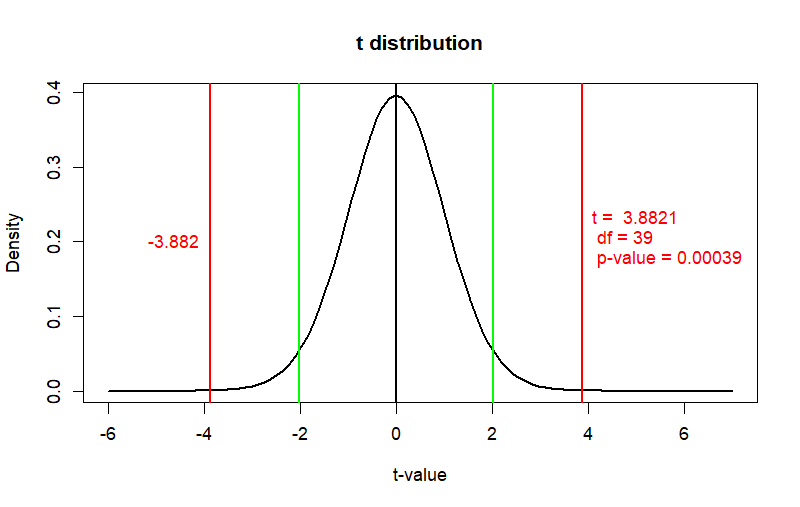
lo2 <- qt(0.05/2,sz-1)
hi2 <- -lo2
lo3 <- qt(0.01/2, sz-1)
hi3 <- -lo3
curve(dt(x, df=sz-1), from = -6, to = 7,
main = "t distribution",
ylab = "Density", xlab = "t-value", col = "black", lwd = 2)
abline(v=0, col="black", lwd=2)
abline(v=c(lo2, hi2), col="green", lwd=2)
abline(v=c(t.cal,-t.cal), col="red", lwd=2)
text(x=-t.cal, y=.2, label=paste("t = ", round(-t.cal,4),
"\n", "df =", sz-1, "\n", "p-value =", round(prob,5)),
col="red", pos=4)
text(x=t.cal, y=.2, label=c(round(t.cal,3)), col="red", pos=2)
cat(t.cal, sz-1, prob)
ro.hypothesis.testing
>
> rm(list=ls())
> rnorm2 <- function(n,mean,sd){ mean+sd*scale(rnorm(n)) }
> ss <- function(x) { sum((x-mean(x))^2) }
> z.test <- function(sam, mu, sigma) {
+ diff <-mean(sam) - mu
+ se <- sigma / sqrt(length(sam))
+ z.cal <- diff / se
+ p.val <- pnorm(abs(z.cal), lower.tail = F)*2
+ two <- abs(qnorm(.05/2))
+ return(cat(
+ " z value:", round(z.cal,5), '\n',
+ "p value:", round(p.val,8), '\n',
+ "diff: ", mean(sam), "-", mu, "=", mean(sam)-mu, '\n',
+ "se: ", se, '\n',
+ "95% CI: ", c(mu-two*se, mu+two*se) ) )
+ }
>
> ################################
> set.seed(1001)
> N.p <- 1000000
> m.p <- 100
> sd.p <- 10
>
> p1 <- rnorm2(N.p, m.p, sd.p)
> p2 <- rnorm2(N.p, m.p+6, sd.p)
>
> ################################
> sz <- 10
> iter <- 100000
> # n = 10 일 때의 p1에 대한 sampling dist 은 아래 시뮬레이션으로
> # 구해볼 수 있다.
> means <- rep(NA, iter)
> for (i in 1:iter) {
+ s1 <- sample(p1, sz, replace = T)
+ means[i] <- mean(s1)
+ }
> mean(means)
[1] 99.9946
> var(means)
[1] 9.95743
> sd(means)
[1] 3.15554
>
> # CLT에 의하면 위이 값은
> m.means <- mean(p1)
> ms.means <- c(var(p1)/sz)
> sd.means <- c(sqrt(var(p1)/sz))
> m.means
[1] 100
> ms.means
[1] 10
> sd.means
[1] 3.162278
>
> # 위의 시뮬레이션으로 구한 sampling dist 대신에
> # 정확한 mean과 se 값을 갖는 집합 sdc를 만든다
> sdc <- rnorm2(iter, m.means, sd.means)
> mean(sdc)
[1] 100
> var(sdc)
[,1]
[1,] 10
> sd(sdc)
[1] 3.162278
>
> zsdc <- scale(sdc)
> m.zsdc <- mean(zsdc)
> ms.zsdc <- var(zsdc)
> sd.zsdc <- sd(zsdc)
> m.zsdc
[1] -2.40102e-17
> ms.zsdc
[,1]
[1,] 1
> sd.zsdc
[1] 1
>
> se2 <- c(sqrt(var(p2)/sz))
> z.p2 <- c((mean(p2)-mean(p1))/se2)
> sdc2 <- rnorm2(iter, mean(p2), se2)
> zsdc2 <- scale(sdc2)+z.p2
> mean(zsdc2)
[1] 1.897367
> sd(zsdc2)
[1] 1
>
> curve(dnorm(x), from = -4, to = z.p2+4,
+ main = "normalized distribution of sample means from p1 and p2",
+ ylab = "Density", xlab = "t-value", col = "black", lwd = 2)
> curve(dnorm(x-(z.p2)), from = z.p2-3, to = z.p2+3, add = T,
+ main = "Distribution Curve",
+ ylab = "Density", xlab = "t-value", col = "blue", lwd = 2, lty=2)
> abline(v=mean(zsdc), col='black', lwd=2)
> abline(v=mean(zsdc2), col='blue', lwd=2)
> mean(zsdc2)
[1] 1.897367
> text(x=mean(zsdc), y=.1, label=paste(round(mean(zsdc),4)), pos=4)
> text(x=mean(zsdc2), y=.1, label=paste(round(mean(zsdc2),4)), pos=4)
>
> #
> lo1 <- qnorm(.32/2)
> hi1 <- -lo1
> lo2 <- qnorm(.05/2)
> hi2 <- -lo2
> lo3 <- qnorm(.01/2)
> hi3 <- -lo3
> c(lo1,hi1)
[1] -0.9944579 0.9944579
> c(lo2,hi2)
[1] -1.959964 1.959964
> c(lo3,hi3)
[1] -2.575829 2.575829
>
> curve(dnorm(x), from = -4, to = 2+4,
+ main = "normalized distribution of sample means from p1",
+ ylab = "Density", xlab = "z-value", col = "black", lwd = 2)
> abline(v=0, col="black", lwd=2)
> abline(v=c(lo1, hi1, lo2, hi2, lo3, hi3),
+ col=c("red","red", "blue", "blue", "orange", "orange"),
+ lwd=2)
> text(x=hi1, y=.2, label=paste(round(hi1,3), "(1)", "\n","86%"), pos=4)
> text(x=hi2, y=.15, label=paste(round(hi2,3),"(2)", "\n","95%"), pos=4)
> text(x=hi3, y=.1, label=paste(round(hi3,3), "(3)", "\n","99%"), pos=4)
>
> mean.of.sample.a <- mean(sdc)+ 1.5*sd(sdc)
> mean.of.sample.a
[1] 104.7434
> diff <- (mean.of.sample.a - mean(sdc))
> se.z <- sd(p1)/sqrt(sz)
> diff
[1] 4.743416
> se.z
[1] 3.162278
> z.score <- diff / se.z
> z.score
[1] 1.5
> prob <- pnorm(z.score, lower.tail = F)*2
> prob
[1] 0.1336144
>
> curve(dnorm(x), from = -4, to = z.p2+4,
+ main = "normalized distribution of sample means from p1 with z score 1.5",
+ ylab = "Density", xlab = "z-value", col = "black", lwd = 2)
> abline(v=0, col="black", lwd=2)
> abline(v=c(lo2, hi2), col=c("blue"), lwd=2)
> abline(v=z.score,col='red', lwd=2)
> abline(v=-z.score, col='red', lwd=2)
> text(x=z.score, y=0.2,
+ label = paste(" z score =", z.score,
+ "\n", "pnorm(-z.score)*2 =", round(prob,5)),
+ pos=4, col='red')
>
> # 새로운 UI로 게임을 하도록 한 후
> # UI점수를 10명에게 구했다고 가정하고
> # 새로운 UI점수가 기존의 p1 paramter와
> # 다른지 테스트 해보라
>
> # z-test 가설검증에 실패하는 경우, 사실은 성공해야 하는데
> # p2의 평균이 106점이고 p1은 100점이니 p2에서 sample을 취
> # 하면 샘플의 평균과 p1의 평균은 다르다고 판단될 것이다.
> # 아래는 그럼에도 불구하고 실패하는 경우이다.
> set.seed(111)
> smp <- sample(p2, sz, replace=T)
> m.smp <- mean(smp)
> m.smp
[1] 104.4742
> diff <- m.smp - mean(p1)
> se.z <- sqrt(var(p1)/sz)
> z.cal1 <- diff / se.z
> prob1 <- pnorm(abs(z.cal1), lower.tail = F)*2
> print(c(z.cal1, sz, prob1))
[1] 1.4148817 10.0000000 0.1571032
> z.test(smp, mean(p1), sd(p1))
z value: 1.41488
p value: 0.1571032
diff: 104.4742 - 100 = 4.474249
se: 3.162278
95% CI: 93.80205 106.198>
> curve(dnorm(x), from = -4, to = z.p2+4,
+ main = "normalized distribution of sample means \n testing with a sample from p2 (failed)",
+ ylab = "Density", xlab = "z-value", col = "black", lwd = 2)
> abline(v=0, col="black", lwd=2)
> abline(v=c(lo2, hi2), col=c("blue"), lwd=2)
> abline(v=z.cal1,col='red', lwd=2)
> abline(v=-z.cal1, col='red', lwd=2)
> text(x=z.cal1, y=0.2,
+ label = paste(" z score =", round(z.cal1,4),
+ "\n", "pnorm(-z.cal1)*2 =", round(prob1,4)),
+ pos=4, col='red')
>
>
> # 같은 방법으로 했는데 성공한 경우
> set.seed(211)
> smp <- sample(p2,sz,replace=T)
> m.smp <- mean(smp)
> m.smp
[1] 110.1154
> diff <- m.smp - mean(p1)
> se.z <- sqrt(var(p1)/sz)
> z.cal2 <- diff / se.z
> prob2 <- pnorm(abs(z.cal2), lower.tail = F)*2
> print(c(z.cal2, sz, prob2))
[1] 3.198763975 10.000000000 0.001380181
> z.test(smp, mean(p1), sd(p1))
z value: 3.19876
p value: 0.00138018
diff: 110.1154 - 100 = 10.11538
se: 3.162278
95% CI: 93.80205 106.198>
> z.p2 <- (mean(p2)-mean(p1))/se2
> z.p2
[1] 1.897367
> curve(dnorm(x), from = -5, to = z.p2+5,
+ main = "normalized distribution of sample means \n testing with a sample from p2 (succeeded)",
+ ylab = "Density", xlab = "z-value", col = "black", lwd = 2)
> abline(v=0, col='black', lwd=2)
> z.cal1
[,1]
[1,] 1.414882
> z.cal2
[,1]
[1,] 3.198764
> two <- qnorm(.05/2)
> two
[1] -1.959964
> abline(v=c(two, -two), col='black', lwd=2)
> abline(v=c(-z.cal2, z.cal2), col='green', lwd=2)
> text(x=z.cal2, y=.3,
+ label=paste("z.cal =", round(z.cal2,4), "\n",
+ "pnorm(-z.cal2)*2 =", "\n", round(prob2,5)),
+ col="darkgreen", cex=1, pos=4)
> text(x=-z.cal2, y=.3,
+ label=paste(round(-z.cal2,4)),
+ col="darkgreen", cex=1, pos=2)
>
>
> # type i and type ii error
> z.p2 <- (mean(p2)-mean(p1))/se2
> z.p2
[1] 1.897367
> curve(dnorm(x), from = -4.7, to = z.p2+4,
+ main = "Distribution Curve",
+ ylab = "Density", xlab = "z-value", col = "black", lwd = 2)
> curve(dnorm(x-(z.p2)), from = z.p2-3, to = z.p2+3, add = T,
+ main = "Distribution Curve",
+ ylab = "Density", xlab = "z-value", col = "blue", lwd = 2, lty=2)
> abline(v=0, col='black', lwd=2)
> z.cal1
[,1]
[1,] 1.414882
> z.cal2
[,1]
[1,] 3.198764
> two <- qnorm(.05/2)
> two
[1] -1.959964
> abline(v=c(two, -two), col='black', lwd=2)
> abline(v=c(-z.cal1, z.cal1), col='red', lwd=2)
> text(x=z.cal1, y=.1,
+ label=paste("z.cal =", round(z.cal1,4), "\n",
+ "pnorm(-z.cal1)*2 =", "\n", round(prob1,3)),
+ col="red", cex=1, pos=4)
> text(x=-z.cal1, y=.1,
+ label=paste(round(-z.cal1,4)),
+ col="red", cex=1, pos=2)
>
> abline(v=c(-z.cal2, z.cal2), col='green', lwd=2)
> text(x=z.cal2, y=.3,
+ label=paste("z.cal =", round(z.cal2,4), "\n",
+ "pnorm(-z.cal2)*2 =", "\n", round(prob2,5)),
+ col="darkgreen", cex=1, pos=4)
> text(x=-z.cal2, y=.3,
+ label=paste(round(-z.cal2,4)),
+ col="darkgreen", cex=1, pos=2)
>
>
> ############################
> # one sample t-test
> ############################
> set.seed(99)
> sz <- 20
> smp <- sample(p2, sz, replace = T)
> m.smp <- mean(smp)
> diff <- m.smp - mean(p1)
> se.z <- sqrt(var(smp)/sz)
> df.smp <- sz - 1
> t.cal <- (diff/se.z)
> prob <- pt(t.cal, df.smp, lower.tail = F)*2
> se.z
[1] 1.809134
> qt(.05/2, df.smp)
[1] -2.093024
> lo2 <- qt(.05/2, df.smp)
> hi2 <- -lo2
> m.smp+lo2*se.z
[1] 102.5239
>
> curve(dt(x, df.smp), from = -6, to = 7,
+ main = "t distribution",
+ ylab = "Density", xlab = "t-value", col = "black", lwd = 2)
> abline(v=0, col="black", lwd=2)
> abline(v=c(lo2, hi2), col="green", lwd=2)
> abline(v=c(-t.cal, t.cal), col="red", lwd=2)
> text(x=t.cal,y=.2,
+ label=paste("t =", round(t.cal,3),
+ "\n", "df =", df.smp,
+ "\n", "p value =", round(prob, 5)),
+ pos = 4, col="red", cex=1)
>
> prob
[1] 0.002460977
>
> print(c(t.cal, df.smp, prob))
[1] 3.488086575 19.000000000 0.002460977
> print(c(m.smp+lo2*se.z, m.smp+hi2*se.z))
[1] 102.5239 110.0970
> cat("t =", t.cal, ", df =", round(df.smp,0), ", p-value =", prob,
+ "\n", "95% confidence interval =", m.smp+lo2*se.z, m.smp+hi2*se.z)
t = 3.488087 , df = 19 , p-value = 0.002460977
95% confidence interval = 102.5239 110.097> t.test(smp, mu=mean(p1))
One Sample t-test
data: smp
t = 3.4881, df = 19, p-value = 0.002461
alternative hypothesis: true mean is not equal to 100
95 percent confidence interval:
102.5239 110.0970
sample estimates:
mean of x
106.3104
>
> #################################
> # t-test 2 group
> #################################
> set.seed(169)
> sz.a <- 25
> sz.b <- 25
> group.a <- sample(p1, sz.a)
> group.b <- sample(p2, sz.b)
> m.a <- mean(group.a)
> m.b <- mean(group.b)
> ss.a <- ss(group.a)
> ss.b <- ss(group.b)
> df.a <- sz.a - 1
> df.b <- sz.b - 1
> df <- df.a+df.b
> ss.a
[1] 2225.751
> ss.b
[1] 2783.816
> df.a
[1] 24
> df.b
[1] 24
>
> pooled.v <- (ss.a+ss.b)/(df.a+df.b)
> pooled.v
[1] 104.366
> se.s <- sqrt(pooled.v/sz.a+pooled.v/sz.b)
> se.s
[1] 2.889512
> diff <- m.a-m.b
> t.cal <- diff/se.s
> t.cal
[1] -3.070212
>
> prob <- pt(abs(t.cal), df=df, lower.tail = F)*2
>
> t.cal
[1] -3.070212
> df
[1] 48
> prob
[1] 0.003515457
>
> t.test(group.a, group.b, var.equal = T)
Two Sample t-test
data: group.a and group.b
t = -3.0702, df = 48, p-value = 0.003515
alternative hypothesis: true difference in means is not equal to 0
95 percent confidence interval:
-14.681167 -3.061661
sample estimates:
mean of x mean of y
101.0286 109.9000
>
> lo2 <- qt(.05/2, df)
> hi2 <- -lo2
> c(lo2, hi2)
[1] -2.010635 2.010635
>
> curve(dt(x, df=df), from = -6, to = 6,
+ main = "t distribution curve",
+ ylab = "Density", xlab = "t-value", col = "black", lwd = 2)
> abline(v=0, col="black", lwd=2)
> abline(v=c(lo2, hi2), col=c("blue"), lwd=2)
> abline(v=c(t.cal, -t.cal), col='red', lwd=2)
> text(x=t.cal, y=0.2,
+ label = paste(" t =", round(t.cal, 4),
+ "\n",
+ "df =", df, "\n",
+ "pt(-t.cal)*2 =", round(prob, 4)),
+ pos=4, col='red')
>
> print(paste(t.cal, df, prob))
[1] "-3.07021182079817 48 0.00351545738746208"
> t.test(group.a, group.b, var.equal = T)
Two Sample t-test
data: group.a and group.b
t = -3.0702, df = 48, p-value = 0.003515
alternative hypothesis: true difference in means is not equal to 0
95 percent confidence interval:
-14.681167 -3.061661
sample estimates:
mean of x mean of y
101.0286 109.9000
> t.cal
[1] -3.070212
> # t.cal=diff/se
> t.cal * se.s
[1] -8.871414
> diff
[1] -8.871414
> diff+lo2*se.s
[1] -14.68117
> diff+hi2*se.s
[1] -3.061661
> (t.cal+lo2)*se.s
[1] -14.68117
> (t.cal+hi2)*se.s
[1] -3.061661
>
> ######################
> # 4번째 케이스 t-test
> ######################
> set.seed(37)
> sz <- 40
> time.a <- sample(p1,sz)
> time.b <- sample(p2,sz)
> diff.time <- time.a-time.b
> m.a <- mean(time.a)
> m.b <- mean(time.b)
> diff <- m.a-m.b
> diff
[1] -8.674375
> se.s <- sd(diff.time)/sqrt(sz)
> t.cal <- diff/se.s
> df <- sz - 1
> prob <- pt(abs(t.cal), df=sz-1, lower.tail = F)*2
> t.cal
[1] -3.88213
> df
[1] 39
> prob
[1] 0.0003888961
> diff
[1] -8.674375
>
> m.diff.time <- mean(diff.time)
> se.s
[1] 2.234437
>
> t.test(time.a, time.b, paired=T)
Paired t-test
data: time.a and time.b
t = -3.8821, df = 39, p-value = 0.0003889
alternative hypothesis: true mean difference is not equal to 0
95 percent confidence interval:
-13.193950 -4.154799
sample estimates:
mean difference
-8.674375
>
> m.diff.time
[1] -8.674375
> se.s
[1] 2.234437
> lo1 <- qt(0.32/2,sz-1)
> hi1 <- -lo1
> lo2 <- qt(0.05/2,sz-1)
> hi2 <- -lo2
> lo3 <- qt(0.01/2, sz-1)
> hi3 <- -lo3
>
> curve(dt(x, df=sz-1), from = -6, to = 7,
+ main = "t distribution",
+ ylab = "Density", xlab = "t-value", col = "black", lwd = 2)
>
> abline(v=0, col="black", lwd=2)
> abline(v=c(lo2, hi2), col="green", lwd=2)
> abline(v=c(t.cal,-t.cal), col="red", lwd=2)
> text(x=-t.cal, y=.2, label=paste("t = ", round(-t.cal,4),
+ "\n", "df =", sz-1, "\n", "p-value =", round(prob,5)),
+ col="red", pos=4)
> text(x=t.cal, y=.2, label=c(round(t.cal,3)), col="red", pos=2)
>
> cat(t.cal, sz-1, prob)
-3.88213 39 0.0003888961
>