Independent sample t-test 의 standard error 값을 구하는 방법이 얼른 머리에 들어오지 않으면,
see mean and variance of the sample mean: 샘플평균들의 집합이 가지는 평균과 분산 (CLT)
sa statistical review: 분산의 계산
두 샘플을 취해서 평균을 구한 후 (mean of A, mean of B), 그 차이를 기록하는 것을 무한히 하여 그 분포를 구하는 것은 아래와 같이 정리, 이해할 수 있다.
\begin{eqnarray}
E \left[ \overline{X} - \overline{Y} \right] & = & E \left[ \overline{X} \right] - E \left[ \overline{Y} \right]
& = & {\mu_{X}} - {\mu_{Y}} \\
V \left[ \overline{X} - \overline{Y} \right] & = & V \left[ \overline{X} \right] + V \left[ \overline{Y} \right]
& = & \displaystyle \dfrac {\sigma^2_{X}} {n_{X}} + \dfrac {\sigma^2_{Y}} {n_{Y}} \\
\end{eqnarray}
- 위가 말하는 것은 X 모집단과 Y 모집단의 평균이 같다면 거기서 뽑은 두 샘플평균의 차이를 모아 놓은 분포는
- 1. Normal distribution이고
- 2. 평균은 두 모집단 평균의 차이가 되고
- 3. 분산은 위의 식처럼 될 것이다.
- 보통의 경우 모집단의 파라미터를 알고 있는 경우는 없다. 따라서 샘플의 분산을 이용해서 SE값을 구한다.
\begin{eqnarray} V \left[ \overline{X} - \overline{Y} \right] & = & V \left[ \overline{X} \right] + V \left[ \overline{Y} \right] & = & \displaystyle \dfrac {s^2_{X}} {n_{X}} + \dfrac {s^2_{Y}} {n_{Y}} \\ \end{eqnarray}
- 샘플의 크기가 다른 경우에는 두 샘플을 함께 고려하여 분산을 구한다.
\begin{eqnarray} V \left[ \overline{X} \right] + V \left[ \overline{Y} \right] & = & \displaystyle \dfrac {s^2_{\text{pooled}}} {n_{X}} + \dfrac {s^2_{\text{pooled}}} {n_{Y}} \\ \text{where } \;\;\; s^2_{\text{pooled}} & = & \frac {SS(X) + SS(Y)} {\text{df}_X + \text{df}_Y} \\ \end{eqnarray}
rs.two.sample.t-test
rm(list=ls())
rnorm2 <- function(n,mean,sd){
mean+sd*scale(rnorm(n))
}
ss <- function(x) {
sum((x-mean(x))^2)
}
N.p <- 1000000
m.p <- 100
sd.p <- 10
set.seed(101)
p1 <- rnorm2(N.p, m.p, sd.p)
mean(p1)
sd(p1)
p2 <- rnorm2(N.p, m.p+10, sd.p)
mean(p2)
sd(p2)
sz1 <- 50
sz2 <- 50
df1 <- sz1 - 1
df2 <- sz2 - 1
df.tot <- df1 + df2
iter <- 100000
mdiffs <- rep(NA, iter)
means.s1 <- rep(NA, iter)
means.s2 <- rep(NA, iter)
tail(mdiffs)
for (i in 1:iter) {
# means <- append(means, mean(sample(p1, s.size, replace = T)))
s1 <- sample(p1, sz1, replace = T)
s2 <- sample(p2, sz2, replace = T)
means.s1[i] <- mean(s1)
means.s2[i] <- mean(s2)
mdiffs[i] <- mean(s1)-mean(s2)
}
# 정리, 증명에 의한 계산
mu <- mean(p1) - mean(p2)
# var(x1bar-x2bar) = var(x1bar) + var(x2bar)
# var(x1bar) = var(x1)/n, n = sample size
ms <- var(p1)/sz1 + var(p2)/sz2
se <- sqrt(ms)
mu
ms
se
# 시뮬레이션에 의한 집합 (distribution)
# mdiffs
m.diff <- mean(mdiffs)
var.diff <- var(mdiffs)
sd.diff <- sd(mdiffs)
m.diff
var.diff
sd.diff
var(means.s1)
var(p1)/sz1
var(means.s2)
var(p2)/sz2
var(means.s1-means.s2)
var(means.s1) + var(means.s2) - 2 * cov(means.s1, means.s2)
# 두 집합이 완전히 독립적일 때 cov = 0 이므로
var(p1)/sz1 + var(p2)/sz2
var.diff <- (var(p1)/sz1) + (var(p2)/sz2)
var.diff
sqrt(var.diff)
se.diff <- sqrt(var.diff)
se.diff
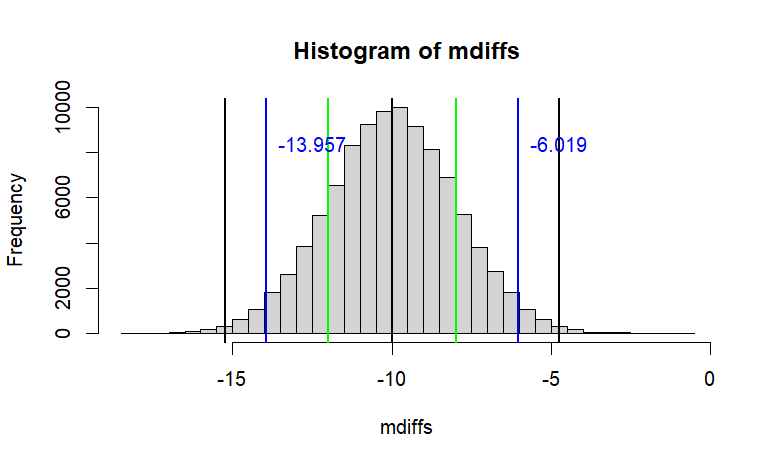
# 이것을 그래프로 그려보면
hist(mdiffs, breaks=50,
main=paste("Histogram of mdiffs",
"\n","with 95% CI (blue)"))
abline(v=mean(mdiffs),
col="black", lwd=2)
se.diff
one <- qt(1-(.32/2), df=df.tot)
two <- qt(1-(.05/2), df=df.tot)
thr <- qt(1-(.01/2), df=df.tot)
ci68 <- se.diff*one
ci95 <- se.diff*two
ci99 <- se.diff*thr
abline(v=c(m.diff-ci68, m.diff-ci95, m.diff-ci99,
m.diff+ci68, m.diff+ci95, m.diff+ci99),
col=c("green", "blue", "black"), lwd=2)
text(x=m.diff-ci95, y=iter/12,
labels=paste(round(m.diff-ci95,3)),
col="blue", pos = 4)
text(x=m.diff+ci95, y=iter/12,
labels=paste(round(m.diff+ci95,3)),
col="blue", pos = 4)
# 위는 이론적으로 생각해 보는 것이고
# 아래는 실제로 2 샘플을 취한 것
s1 <- sample(p1, sz1, replace = T)
s2 <- sample(p2, sz2, replace = T)
m.diff <- mean(s1)-mean(s2)
m.diff # <- 100 번 중 95번은 위의 블루라인 안쪽에서
# 이 때의 pooled variance는 ss(s1)을 이용해서 구함
# population의 parameter는 모르는 상태로 가정
pv <- (ss(s1) + ss(s2))/(df1 + df2)
pv
ms1 <- ss(s1)/df1
ms2 <- ss(s2)/df2
ms1
ms2
(ms1 + ms2)/2
# se <- sqrt(ms1/sz1 + ms2/sz2)
# se
se.z <- sqrt(pv/sz1 + pv/sz2)
se.z
diff <- mean(s1)-mean(s2)
t.cal <- diff / se.z
t.cal
df.tot
p.val <- pt(abs(t.cal), df.tot, lower.tail = F)*2
mean.diff <- mean(s1)-mean(s2)
# two <- -2 에 해당하는 정확한 값
two <- qt(.05/2, df.tot)
lo2 <- se.z * two
lo2
paste(c(t.cal, df.tot, p.val))
mean.diff + c(lo2, -lo2)
t.test(s1,s2, var.equal = T)
zdiffs <- scale(mdiffs)
se.diff <- sd.diff
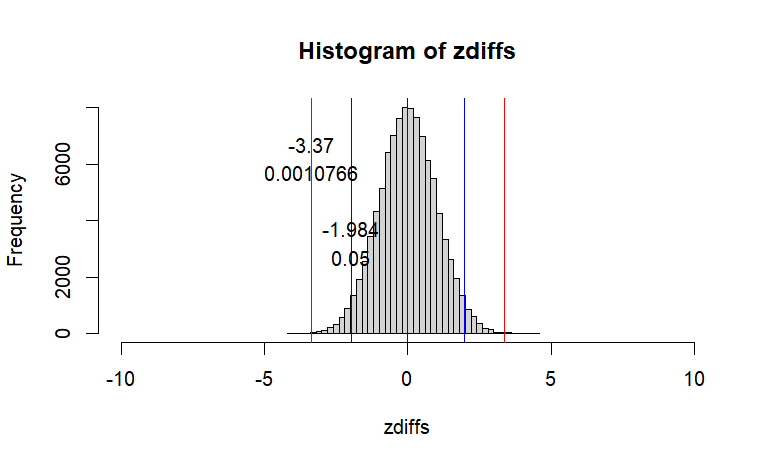
hist(zdiffs, breaks=50,
xlim =c(-10,10))
two
abline(v=c(0, 0-two, 0+two), col="blue")
text(x=two,y=iter*.03,labels=round(two,3), pos=3)
text(x=two,y=iter*.02, labels=.05,pos=3)
abline(v=c(t.cal,-t.cal), col="red")
text(x=t.cal,y=iter*.06,labels=round(t.cal,3),pos=3)
text(x=t.cal, y=iter*.05,labels=round(p.val,7),pos=3)
p.val
###
# what if s1 and s2 are from
# the same pop?
# the distribution of sample mean
# difference should be
# normal, mu = 0, var = var.p1/s.size + var.p2/s.size
iter <- 100000
# means <- c()
mdiffs <- rep(NA, iter)
means.s3 <- rep(NA, iter)
means.s4 <- rep(NA, iter)
tail(mdiffs)
for (i in 1:iter) {
# means <- append(means, mean(sample(p1, s.size, replace = T)))
s3 <- sample(p1, sz1, replace = T)
s4 <- sample(p1, sz2, replace = T)
means.s3[i] <- mean(s3)
means.s4[i] <- mean(s4)
mdiffs[i] <- mean(s3)-mean(s4)
}
mu <- mean(p1) - mean(p1)
ms <- var(p1)/sz1 + var(p1)/sz2
se <- sqrt(ms)
mu
ms
se
m.diff <- mean(mdiffs)
var.diff <- var(mdiffs)
sd.diff <- sd(mdiffs)
m.diff
var.diff
sd.diff
s3 <- sample(p1, sz1, replace=T)
s4 <- sample(p1, sz2, replace=T)
t.test(s3, s4, var.equal=T)
print(m.diff <- mean(s3)-mean(s4))
# 위의 value는 0을 중심으로 -4 +4 사이에 있을 확률이 95퍼센트이다.
# 정확히는 mean difference = p1 - p1 = 0 se = 2 95% Interval = two
print(two <- qt(.05/2,df.tot))
# 아래에 ss(p1) 대신에 ss(s3)를 사용한 것은
# 모집단의 정보가 없음을 가정하기 때문에
pv <- (ss(s3)+ss(s4))/(df1+df2)
se.diff <- sqrt(pv/sz1 + pv/sz2)
se.diff
lo <- two*se.diff
hi <- -lo
c(lo, hi)
# let's see
# but, this is exactly what distribution of sample differences
# in the first place
iter <- 1000
means.s3 <- means.s4 <- rep(NA, iter)
mdiffs <- rep(NA, iter)
for (i in 1:iter) {
# means <- append(means, mean(sample(p1, s.size, replace = T)))
s3 <- sample(p1, sz1, replace = T)
s4 <- sample(p1, sz2, replace = T)
means.s3[i] <- mean(s3)
means.s4[i] <- mean(s4)
mdiffs[i] <- mean(s3)-mean(s4)
}
table(mdiffs < -4 | mdiffs > 4)
ro.two.sample.t-test
>
>
> rm(list=ls())
> rnorm2 <- function(n,mean,sd){
+ mean+sd*scale(rnorm(n))
+ }
> ss <- function(x) {
+ sum((x-mean(x))^2)
+ }
>
> N.p <- 1000000
> m.p <- 100
> sd.p <- 10
>
> set.seed(101)
> p1 <- rnorm2(N.p, m.p, sd.p)
> mean(p1)
[1] 100
> sd(p1)
[1] 10
>
> p2 <- rnorm2(N.p, m.p+10, sd.p)
> mean(p2)
[1] 110
> sd(p2)
[1] 10
>
> sz1 <- 50
> sz2 <- 50
> df1 <- sz1 - 1
> df2 <- sz2 - 1
> df.tot <- df1 + df2
>
> iter <- 100000
> mdiffs <- rep(NA, iter)
> means.s1 <- rep(NA, iter)
> means.s2 <- rep(NA, iter)
> tail(mdiffs)
[1] NA NA NA NA NA NA
>
> for (i in 1:iter) {
+ # means <- append(means, mean(sample(p1, s.size, replace = T)))
+ s1 <- sample(p1, sz1, replace = T)
+ s2 <- sample(p2, sz2, replace = T)
+ means.s1[i] <- mean(s1)
+ means.s2[i] <- mean(s2)
+ mdiffs[i] <- mean(s1)-mean(s2)
+ }
> # 정리, 증명에 의한 계산
> mu <- mean(p1) - mean(p2)
> # var(x1bar-x2bar) = var(x1bar) + var(x2bar)
> # var(x1bar) = var(x1)/n, n = sample size
> ms <- var(p1)/sz1 + var(p2)/sz2
> se <- sqrt(ms)
>
> mu
[1] -10
> ms
[,1]
[1,] 4
> se
[,1]
[1,] 2
>
> # 시뮬레이션에 의한 집합 (distribution)
> # mdiffs
> m.diff <- mean(mdiffs)
> var.diff <- var(mdiffs)
> sd.diff <- sd(mdiffs)
> m.diff
[1] -9.988058
> var.diff
[1] 4.023723
> sd.diff
[1] 2.005922
>
> var(means.s1)
[1] 2.002125
> var(p1)/sz1
[,1]
[1,] 2
> var(means.s2)
[1] 2.014368
> var(p2)/sz2
[,1]
[1,] 2
> var(means.s1-means.s2)
[1] 4.023723
> var(means.s1) + var(means.s2) - 2 * cov(means.s1, means.s2)
[1] 4.023723
> # 두 집합이 완전히 독립적일 때 cov = 0 이므로
> var(p1)/sz1 + var(p2)/sz2
[,1]
[1,] 4
>
> var.diff <- (var(p1)/sz1) + (var(p2)/sz2)
> var.diff
[,1]
[1,] 4
> sqrt(var.diff)
[,1]
[1,] 2
> se.diff <- sqrt(var.diff)
> se.diff
[,1]
[1,] 2
>
> # 이것을 그래프로 그려보면
> hist(mdiffs, breaks=50,
+ main=paste("Histogram of mdiffs",
+ "\n","with 95% CI (blue)"))
> abline(v=mean(mdiffs),
+ col="black", lwd=2)
>
> se.diff
[,1]
[1,] 2
> one <- qt(1-(.32/2), df=df.tot)
> two <- qt(1-(.05/2), df=df.tot)
> thr <- qt(1-(.01/2), df=df.tot)
>
> ci68 <- se.diff*one
> ci95 <- se.diff*two
> ci99 <- se.diff*thr
>
> abline(v=c(m.diff-ci68, m.diff-ci95, m.diff-ci99,
+ m.diff+ci68, m.diff+ci95, m.diff+ci99),
+ col=c("green", "blue", "black"), lwd=2)
> text(x=m.diff-ci95, y=iter/12,
+ labels=paste(round(m.diff-ci95,3)),
+ col="blue", pos = 4)
> text(x=m.diff+ci95, y=iter/12,
+ labels=paste(round(m.diff+ci95,3)),
+ col="blue", pos = 4)
>
>
> # 위는 이론적으로 생각해 보는 것이고
> # 아래는 실제로 2 샘플을 취한 것
> s1 <- sample(p1, sz1, replace = T)
> s2 <- sample(p2, sz2, replace = T)
> m.diff <- mean(s1)-mean(s2)
> m.diff # <- 100 번 중 95번은 위의 블루라인 안쪽에서
[1] -6.959851
>
> # 이 때의 pooled variance는 ss(s1)을 이용해서 구함
> # population의 parameter는 모르는 상태로 가정
> pv <- (ss(s1) + ss(s2))/(df1 + df2)
> pv
[1] 106.6359
>
> ms1 <- ss(s1)/df1
> ms2 <- ss(s2)/df2
> ms1
[1] 105.3544
> ms2
[1] 107.9175
> (ms1 + ms2)/2
[1] 106.6359
>
> # se <- sqrt(ms1/sz1 + ms2/sz2)
> # se
> se.z <- sqrt(pv/sz1 + pv/sz2)
> se.z
[1] 2.065293
>
> diff <- mean(s1)-mean(s2)
> t.cal <- diff / se.z
>
>
> t.cal
[1] -3.369909
> df.tot
[1] 98
> p.val <- pt(abs(t.cal), df.tot, lower.tail = F)*2
> mean.diff <- mean(s1)-mean(s2)
> # two <- -2 에 해당하는 정확한 값
> two <- qt(.05/2, df.tot)
> lo2 <- se.z * two
> lo2
[1] -4.098508
>
> paste(c(t.cal, df.tot, p.val))
[1] "-3.36990898869323" "98" "0.00107663367234877"
> mean.diff + c(lo2, -lo2)
[1] -11.058359 -2.861343
>
> t.test(s1,s2, var.equal = T)
Two Sample t-test
data: s1 and s2
t = -3.3699, df = 98, p-value = 0.001077
alternative hypothesis: true difference in means is not equal to 0
95 percent confidence interval:
-11.058359 -2.861343
sample estimates:
mean of x mean of y
101.6366 108.5965
>
>
> zdiffs <- scale(mdiffs)
> se.diff <- sd.diff
> hist(zdiffs, breaks=50,
+ xlim =c(-10,10))
> two
[1] -1.984467
> abline(v=c(0, 0-two, 0+two), col="blue")
> text(x=two,y=iter*.03,labels=round(two,3), pos=3)
> text(x=two,y=iter*.02, labels=.05,pos=3)
> abline(v=c(t.cal,-t.cal), col="red")
> text(x=t.cal,y=iter*.06,labels=round(t.cal,3),pos=3)
> text(x=t.cal, y=iter*.05,labels=round(p.val,7),pos=3)
> p.val
[1] 0.001076634
>
> ###
> # what if s1 and s2 are from
> # the same pop?
> # the distribution of sample mean
> # difference should be
> # normal, mu = 0, var = var.p1/s.size + var.p2/s.size
>
> iter <- 100000
> # means <- c()
> mdiffs <- rep(NA, iter)
> means.s3 <- rep(NA, iter)
> means.s4 <- rep(NA, iter)
> tail(mdiffs)
[1] NA NA NA NA NA NA
>
> for (i in 1:iter) {
+ # means <- append(means, mean(sample(p1, s.size, replace = T)))
+ s3 <- sample(p1, sz1, replace = T)
+ s4 <- sample(p1, sz2, replace = T)
+ means.s3[i] <- mean(s3)
+ means.s4[i] <- mean(s4)
+ mdiffs[i] <- mean(s3)-mean(s4)
+ }
>
> mu <- mean(p1) - mean(p1)
> ms <- var(p1)/sz1 + var(p1)/sz2
> se <- sqrt(ms)
>
> mu
[1] 0
> ms
[,1]
[1,] 4
> se
[,1]
[1,] 2
>
> m.diff <- mean(mdiffs)
> var.diff <- var(mdiffs)
> sd.diff <- sd(mdiffs)
> m.diff
[1] -0.00273072
> var.diff
[1] 3.997207
> sd.diff
[1] 1.999302
>
> s3 <- sample(p1, sz1, replace=T)
> s4 <- sample(p1, sz2, replace=T)
> t.test(s3, s4, var.equal=T)
Two Sample t-test
data: s3 and s4
t = 1.7165, df = 98, p-value = 0.08924
alternative hypothesis: true difference in means is not equal to 0
95 percent confidence interval:
-0.5535058 7.6433427
sample estimates:
mean of x mean of y
103.04431 99.49939
> print(m.diff <- mean(s3)-mean(s4))
[1] 3.544918
> # 위의 value는 0을 중심으로 -4 +4 사이에 있을 확률이 95퍼센트이다.
> # 정확히는 mean difference = p1 - p1 = 0 se = 2 95% Interval = two
> print(two <- qt(.05/2,df.tot))
[1] -1.984467
> # 아래에 ss(p1) 대신에 ss(s3)를 사용한 것은
> # 모집단의 정보가 없음을 가정하기 때문에
> pv <- (ss(s3)+ss(s4))/(df1+df2)
> se.diff <- sqrt(pv/sz1 + pv/sz2)
> se.diff
[1] 2.065251
> lo <- two*se.diff
> hi <- -lo
> c(lo, hi)
[1] -4.098424 4.098424
>
> # let's see
> # but, this is exactly what distribution of sample differences
> # in the first place
> iter <- 1000
> means.s3 <- means.s4 <- rep(NA, iter)
> mdiffs <- rep(NA, iter)
>
> for (i in 1:iter) {
+ # means <- append(means, mean(sample(p1, s.size, replace = T)))
+ s3 <- sample(p1, sz1, replace = T)
+ s4 <- sample(p1, sz2, replace = T)
+ means.s3[i] <- mean(s3)
+ means.s4[i] <- mean(s4)
+ mdiffs[i] <- mean(s3)-mean(s4)
+ }
>
> table(mdiffs < -4 | mdiffs > 4)
FALSE TRUE
951 49
>